Why a turtle? Because years ago Bernie Agranoff realised that myo-inositol in its usual conformation in solution (a chair configuration with five hydroxyls equatorial, and the number 2 hydroxyl axial) superficially resembles a turtle. It is easy to avoid any confusion in using the D-numbering now universally employed in the biological literature, simply by remembering that the turtle’s front right flipper is number 1, and his head number 2. The technical term for this is turtle recall.
Inositides and cellular function
Inositides are a diverse and multifunctional group of cellular components, which have in common that myo-inositol is a part of their chemical structure. Such is the diversity of compounds into which evolution has placed inositol that our studies on their metabolism and function touches on almost all aspects of cellular processes. Inositides can be classified into two very different basic types, whose metabolism and functions are mostly separate: the inositol lipids and inositol phosphates. Our work currently focuses on several aspects of both types.
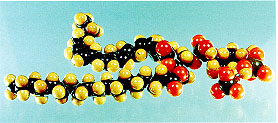 (1) Inositol lipids. Central to most inositide metabolism and function is the inositol lipid, phosphatidylinositol 4,5-bisphosphate (PIP2; see figure). PIP2 is the precursor for at least three second messengers, inositol 1,4,5-trisphosphate (IP3), diacylglycerol, and phosphatidylinositol 3,4,5-trisphosphate (PIP3), and itself plays an important regulatory role in, for example, cytoskeletal function. Thus, the family of enzymes that synthesise it (the phosphatidylinositol phosphate kinases, which we know as PIPkins), will be diverse and under several complex mechanisms of regulation, to ensure appropriate control of PIP2 levels. There are two Types of PIPkin that synthesise PIP2 [1]: the Type I enzymes are PI4P 5-kinases, and catalyse the major route of PIP2 synthesis, while the Type II enzymes are PI5P 4-kinases, which can thus catalyse a minor route of PIP2 synthesis, though the current opinion in the field is that their principal function is to remove (and this regulate the levels of) their substrate PI5P.
(1) Inositol lipids. Central to most inositide metabolism and function is the inositol lipid, phosphatidylinositol 4,5-bisphosphate (PIP2; see figure). PIP2 is the precursor for at least three second messengers, inositol 1,4,5-trisphosphate (IP3), diacylglycerol, and phosphatidylinositol 3,4,5-trisphosphate (PIP3), and itself plays an important regulatory role in, for example, cytoskeletal function. Thus, the family of enzymes that synthesise it (the phosphatidylinositol phosphate kinases, which we know as PIPkins), will be diverse and under several complex mechanisms of regulation, to ensure appropriate control of PIP2 levels. There are two Types of PIPkin that synthesise PIP2 [1]: the Type I enzymes are PI4P 5-kinases, and catalyse the major route of PIP2 synthesis, while the Type II enzymes are PI5P 4-kinases, which can thus catalyse a minor route of PIP2 synthesis, though the current opinion in the field is that their principal function is to remove (and this regulate the levels of) their substrate PI5P.
Our work on Type I PIPkins focuses primarily on the various splice variants of the Type I γ isoform and their physiological function [2,3,23], and with the Type II PIPkins we study all three isoforms, α, β, and γ, and how they are localised and regulated [4,5,24,27, 28], as well as investigating PI5P itself [6,7]. We have recently additionally focused our attention on the quantification of PI4P and PIP2 in cells [29]. Mostly these studies focus on the cytoplasm, but we retain a long-standing interest in the separate, and less well understood, functions of inositol lipids and phosphates within the nucleus [8-10,24].
(2) Inositol phosphates. Once IP3 is generated in the cytosol (where it mobilises calcium), it is rapidly metabolised by two routes: by dephosphorylation (ultimately to inositol) and by phosphorylation to inositol 1,3,4,5-tetrakisphosphate (IP4). Our longest-standing project focuses on the regulation and function of the enzymes that catalyse the latter reaction, the IP3 3-kinases, which includes the possible functions of their product, IP4 [11,12]. Currently we have an active interest in IP3 3-kinase A (the neuronal isoform) [13,14,21] and the more ubiquitous B isoform [15], and how the regulation of their localisation relates to function [25].
We also have a wider interest in other inositol phosphates’ metabolism and function [16], principally focusing on the predominant (by mass) inositol phosphate in eukaryotic cells, IP6 [17,18,22,26], as well having interests in various effectors of both the inositol lipids and phosphates [19]. Recently we have also addressed the detailed kinetics of interaction between PIP2 or PIP3 and proteins whose localisation is regulated by these lipids [20, 30].
